That’s right. If you say something bold, such as “H7N9 vaccines will not be immunogenic“, at least you should back it up with high quality immunoinformatics data. Don’t you think?
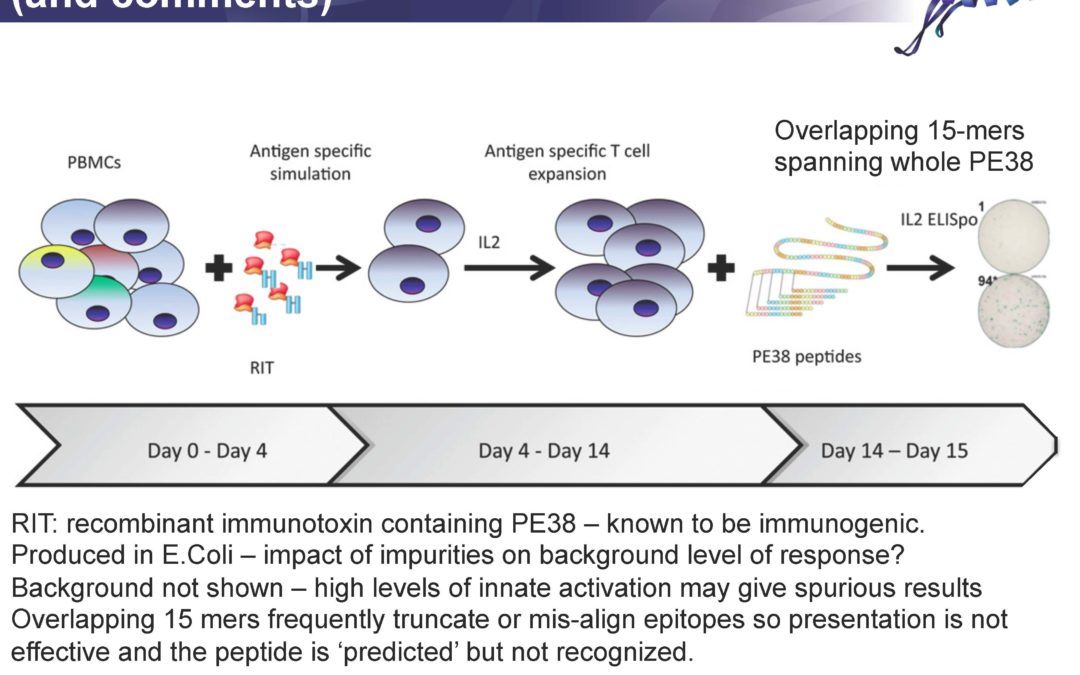
Apparently, some people don’t think so. Take this recent publication for example: The article’s title, states this: Poor correlation between T-cell activation assays and HLA-DR binding prediction algorithms in an immunogenic fragment of Pseudomonas exotoxin A. While that might be the opinion of the authors, careful review of their data would suggest otherwise.
To see a more detailed analysis of PE38 performed using EpiMatrix and correlation with Mazor et, please see this link: Re-analysis of the Mazor/Pastan PE 38 Publication.
And, rather than move the field forward, publishing papers with titles this one, in effect undermines recent, important advances in immunoinformatics that have the potential to move the field forward, leading to more effective, safer, biologics and vaccines. It should come as no surprise that I have a few issues with the title and the paper. Here’s the skinny:
- Sorry folks, IEDB ≠ ISPRI. If I were a co-author on this paper, I would be careful to specify, in my disparaging title, the name of the actual tool that was used, rather than taking a very broad brush and painting the entire field of immunoinformatics black. As in : Poor correlation between T-cell activation assays and IEDB HLA-DR binding prediction algorithms ….
- The tools that were used are not designed for biologics. See our video about that here.
- For example, if you are looking for promiscuous epitopes in outbred populations, look for clusters. Especially if you want to be sure that the peptide will be immunogenic in an outbred population – more useful for deimmunization.
- If you look for clusters, make sure that they aren’t Treg epitopes (a.k.a Tregitopes). As the FDA says in their “Immunogenicity guidelines“, when de-immunizing, Tregitopes should not be removed.
- Etc., etc., etc.
I do have a few positive things to say about the work of Pastan and Mazor. Their publications clearly illustrate how T cell epitope de-immunization can be used to improve the efficacy of biological therapeutics. (As does our work on de-immunization of Botox and FVIII). De-immunization is an under-appreciated discipline. Its application to the development of new and useful medicine has gone largely unexploited, due in part to “opinions” like this one, that immunoinformatics tools are not useful. But – seriously? Can you imagine testing every overlapping peptide in every biologic to determine which peptide to deimmunize? Why do that, when fast, accurate tools already exist?
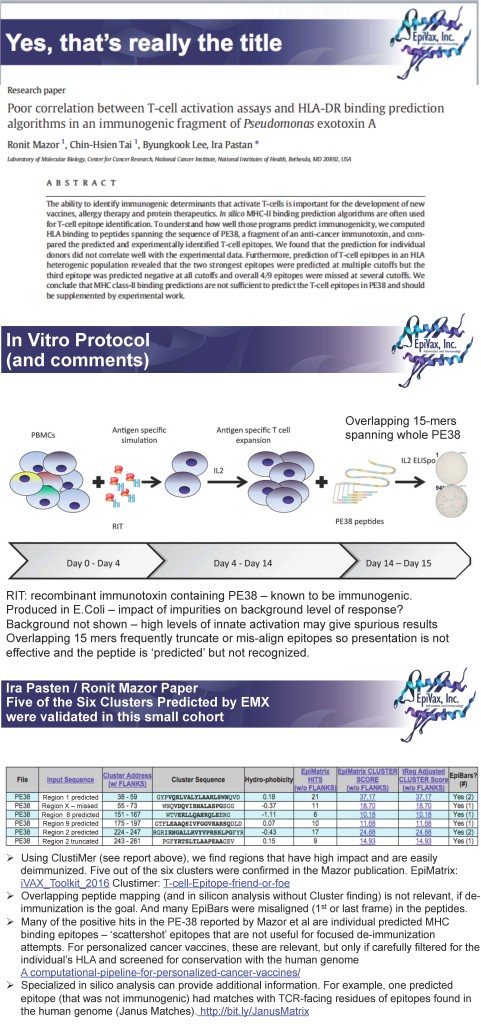 That said, protein deimmunization is not a new idea and this paper doesn’t contribute new perspectives in the field. If anything, the techniques used are quite conservative. In fact, one could say, that using the overlapping peptide approach to mapping biologics for deimmunization is a step back. Perhaps we should make that the title of a forthcoming publication?
That said, protein deimmunization is not a new idea and this paper doesn’t contribute new perspectives in the field. If anything, the techniques used are quite conservative. In fact, one could say, that using the overlapping peptide approach to mapping biologics for deimmunization is a step back. Perhaps we should make that the title of a forthcoming publication?
We believe in the power of immunoinformatics. We believe that immune-engineering using Immunoinformatics tools, like the ones we have been working to develop for the last 18 years – yes, the ones that are currently in commercial use by biologics and vaccine developers all over the world- is the way forward – for biologics and vaccines.
Again, to see a more detailed analysis of PE38 performed using EpiMatrix and correlation with Mazor et, please see this link: Re-analysis of the Mazor/Pastan PE 38 Publication.