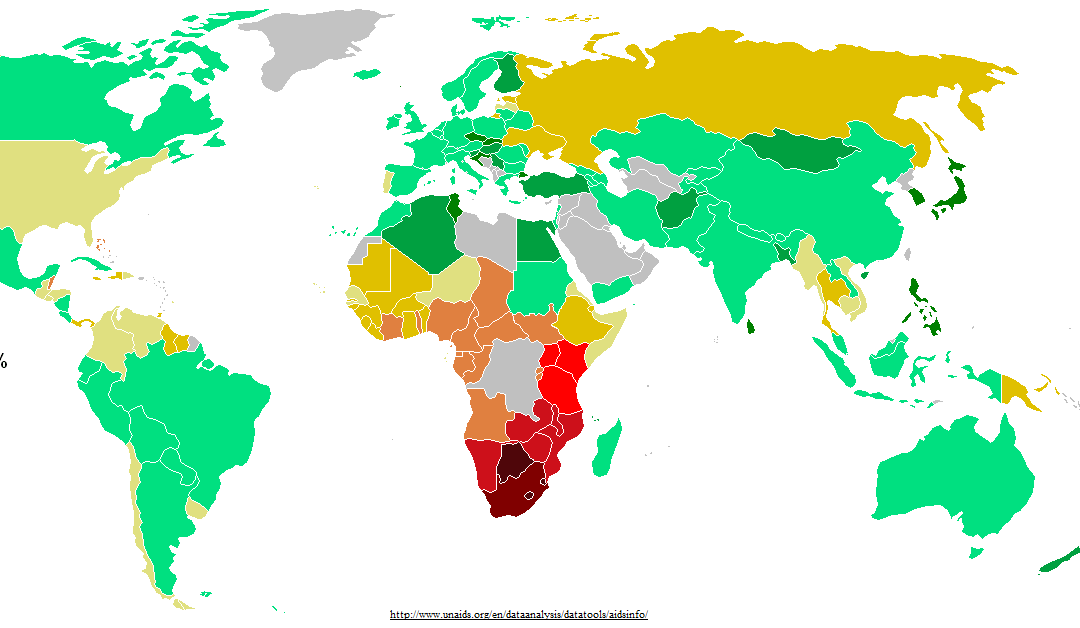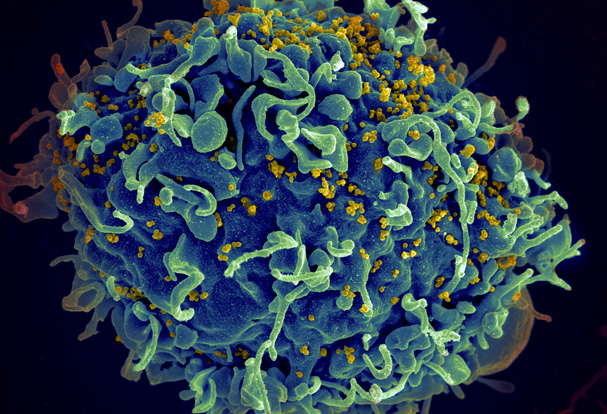

Designing HIV-1 vaccines to reflect viral diversity and the global context of HIV/AIDS
AIDScience 1(2) June 22, 2001.
Bioinformatics tools for identifying class I-restricted epitopes
Methods (Epitope Mapping Issue). Bill Kwok, editor, Methods 29 (2003) 289–298.
Greater CD8+ TCR heterogeneity and functional flexibility in HIV-2 compared to HIV-1 infection
J Immunol. 2003 Jul 1; 171(1):307-16.