ISPRI™
Interactive Screening and Protein Reengineering InterfaceBuild in silico immunogenicity screening into your preclinical R&D process.
The ISPRI™ toolkit by EpiVax is a powerful asset for those developing biologic therapeutics, offering a cutting-edge solution to address challenges in immunogenicity screening and optimization. ISPRI™ is a comprehensive in silico toolkit, available through software licensing agreements, that allows companies to seamlessly integrate immunogenicity screening into their preclinical R&D workflow. The toolkit enables navigation through large lists of ‘hits’, guiding the downselection process by prioritizing leads based on immunogenicity as a key quality attribute. ISPRI™’s modern interface delivers real-time results, even in high-throughput screening scenarios involving hundreds to thousands of sequences simultaneously.
The ability to understand both the overall and regional immunogenic potential of lead candidates, coupled with the capacity to deimmunize or humanize sequences, positions developers to proactively manage and mitigate potential challenges. With ISPRI™, you’re able to strategically direct R&D efforts, leveraging in silico immunogenicity data to inform in vitro and in vivo immunogenicity studies. The toolkit thus becomes a catalyst for enhancing the number of viable leads available for development, ultimately bolstering your R&D pipeline and setting it apart from the rest.
What can ISPRI™ do for you?
-
Set the stage with a high-level assessment of sequence risk potential with whole biologic-level scores driven by T cell epitope mapping in a global assessment.
-
Rank the overall immunogenic potential of each sequence on a normalized Immunogenicity Scale against similar products with known immunogenicity for easy benchmarking.
-
Dig deeper into sequence analysis with a regional assessment that can uncover sequence hotspots that could be key regions for further investigation and optimization, if needed.
-
Gain insight into potential tolerance of key regions by comparing them to human epitopes in an advanced homology assessment.
-
Securely cross-reference your analyses to published data to better enable your decision making.
-
De-risk sequences, if necessary, by strategically introducing point mutations to reduce sequence hotspots.

What’s in the toolkit?
EpiMatrix
What: Identifies putative effector and regulatory T cell epitopes in your sequence.
How: Determines HLA binding potential (a prerequisite for T cell activation) and identifies known regulatory T cell epitopes (Tregitopes) against key HLA supertype alleles across a sequence.
Why: More accurately predict the immunogenic potential of the whole biologic and rank candidates against each other and other biologics with known immunogenicity.
ClustiMer
What: Identifies T cell epitope hotspots in your sequence.
How: Screens sequence for regions of T cell epitope density (short peptides predicted to bind to a large number of HLA alleles).
Why: Dominant T cell epitope clusters can have the potential to drive immune responses in many people, enabling significant immune responses to proteins with low global scores. These areas are key to identify to better understand sequence risk and make excellent targets for deimmunization.
JanusMatrix
What: Analyzes tolerogenic potential of putative T cell epitope content.
How: Assesses T cell epitope cross-conservation with the human proteome by analyzing TCR-facing residues.
Why: Characterizing the depth of sequence T cell epitope content cross-conservation with human epitopes provides key insight to assess the likelihood epitopes may be recognized as self. Epitopes recognized as ‘self’ may be tolerated by or have tolerogenic effects on the immune system. This characterization is key to a more accurate risk assessment.
Homology Analysis
What: Performs basic homology analyses of T cell epitope clusters against human and epitope databases.
How: Cross-references key sequence regions against IEDB and GenBank while keeping your sequences secure.
Why: Leverage published data to further inform your analysis and potential deimmunization choices.
OptiMatrix Premium add-on
What: Deimmunizes high scoring, non-human clusters to reduce immunogenicity.
How: Selects amino acid substitutions that reduce HLA binding potential in key hotspot regions.
Why: Reduce sequence risk by removing putative T cell epitope content.
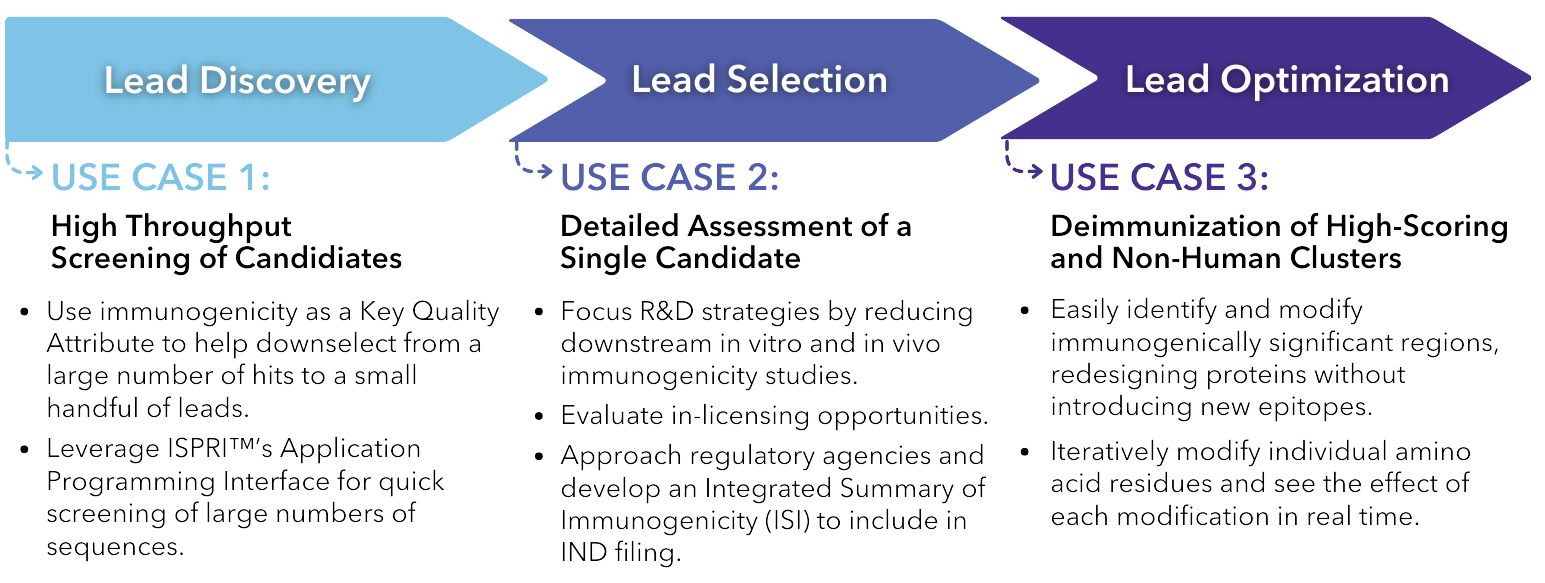
ISPRI™ Use Cases