It’s a new year and welcome to it! Pundits are making a lot of predictions about the decline of science and technology under the new administration in Washington DC, and I’m here with a contrarian view. We bet on biotechnology! We predict that the next four years will see a revolution and acceleration of innovation, regardless of the leadership in The White House.
Why so? Well, because of computers. What? – Of course everyone has a computer! What I mean is that computational immunology is coming of age, and it is clear that proper computational analysis of new drug candidates reduces the time, effort and materials and risk! while producing the best possible biologics for the clinic.
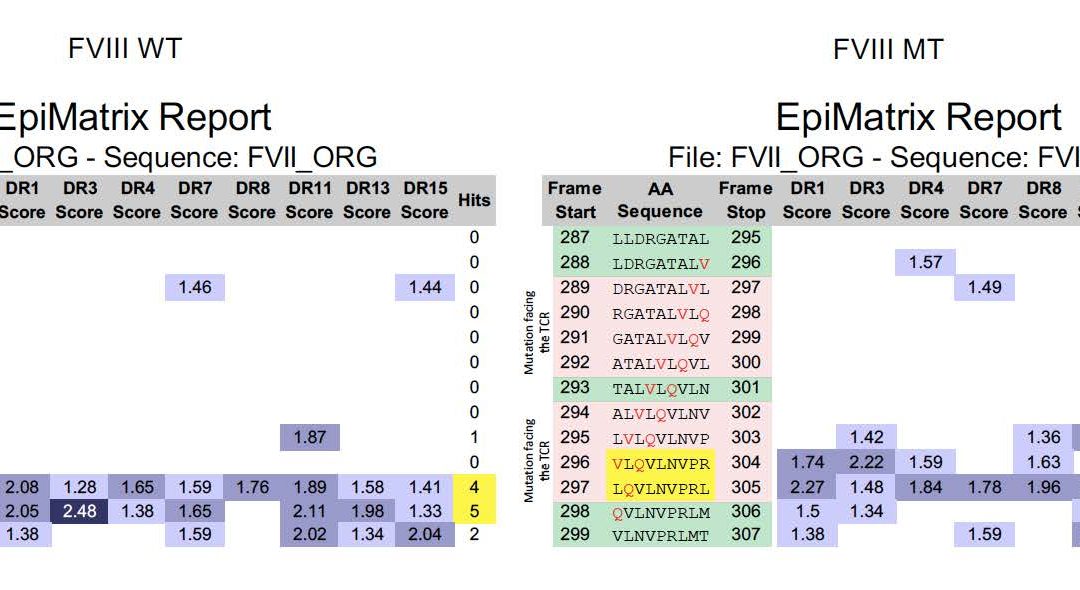
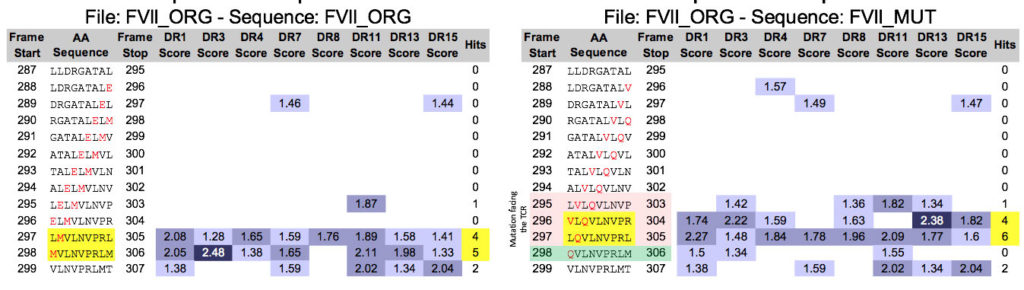
Let’s take a gander at a recent publication by Kasper Lambeth et al, for example. This is a great example – in silico tools were not apparently applied to the mutated FVIIa produced by Novo Nordisk. Now, five years from the time that they scuttled their drug due to immunogenicity (see “More Pesky Anti-Drug-Antibodies, now from Novo Nordisk”) Kasper has collaborated with Zuben Sauna of the FDA to publish a “post hoc” explanation for the problem in Science Translational Medicine (congrats Kasper). As illustrated below, it would have been pretty easy to figure out in advance that the FVIIa re-engineering introduced an immunogenic T cell epitope that was responsible for the ADA.
On the left, we see the original FVIIa sequence and an existing, possibly tolerated epitope. On the right, EpiMatrix and JanusMatrix identify new epitope (highlighted in pink, promiscuity highlighted in yellow) that are substantially different at the T cell face, and the removal of a pre-existing potentially tolerogenic epitope (in green) suggesting that the modification might not be tolerated and could generate an immune response, across HLA types.
In fact, it took all of 5 minutes to upload the modified FVIIa sequence and pinpoint the same risk factor in the recombinant protein with ISPRI. Why do I know? because I ran this analysis on my laptop during one of Kasper’s recent presentations of the data. By the time he was done presenting the data, it was clear what was going on.
5 minutes. What’s better than that?
In fact, yes, there is something better; better than post hoc. How about immune engineering better biologics up front? Yes we can!
We did it with Botox. And we did it with FVIII. And now we did it with Interferon.
Want to know more? Take a look at our recent paper on the development of a DeFT (De-immunized and Functional Therapeutic) version of alpha interferon, authored by Eduardo Mufarrege from Argentina. Eduardo learned how to use the ISPRI toolkit and OptiMatrix to identify regions in the alpha-IFN protein that could be modified and successfully produced a modified version that had equivalent functionality and lower immunogenicity, on an academic budget! Truth be told, the work took a bit longer because he spent a few months at the FDA, but still – 5 years from start to finish, on an academic budget, and with only one part time effort, that’s impressive.
Aha – so that’s DeFT! The strategy of substituting one amino acid for another to make a better biologic. Not so daft after all!
What’s the take home message? Computational Immunology has come of age. Go DeFT, don’t regret! There is really no need to wait for bad news to happen in a clinical trial.
Rather than get all post hoc on us, in 2017, let’s immune engineer! Put your fingers on the keyboard (or send a message in a bottle) and let us know that you want to know more!