When two patients died* due to cardiac complications following treatment with an immunotherapy (T cells armed with a chimeric antigen receptor for a Mage A3 peptide), researchers checked the target peptide again. Using BLAST, there was no obvious cause: no cross-reactivity with any human protein was detected. But where BLAST failed, a new algorithm was able to clarify the relationship between the Mage A3 epitope (EVDPIGHLY) and a class-I restricted T cell epitope found in Titin, a key cardiac protein (ESDPIVAQY, see figure below). TCR facing residues of the two peptides are 100% conserved, potentially explaining the off-target effects.
This relationship was discovered within minutes, using JanusMatrix, a new tool that searches for peptides that have similar MHC binding propensities and homology at the TCR-facing amino acids, within any target genome of interest (1).
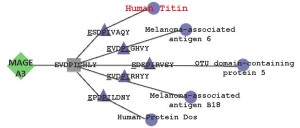
Cytoscape diagram illustrating human genome epitopes containing of cross-conserved TCR-facing residues in the Mage A3 epitope targeted by Chimeric Antigen Receptor (CAR)-bearing T cells in a clinical trial that resulted in two deaths.
JanusMatrix can be used to identify sequences that activate inflammatory responses (T effector epitopes) and other sequences that trigger regulatory T cell (Treg) responses.
Using JanusMatrix, we are able to parse TCR-facing amino acid residues of epitopes and compare them across genomes, to similar-HLA-binding peptides (homologs) derived from self, commensals, or pathogens. Peptides that are conserved with a high number of human homologs can be Tregs (2-5). In contrast, epitopes with low cross-reactivity across the human genome, may drive anti-self immune responses. This may be critically important for immunotherapy in cancer.
Can JanusMatrix also explain off-target effects of other vaccines? We think so. This very same tool is turning up regulatory T cell epitopes in cancer and infectious disease vaccine antigens, that could explain poor antibody seroconversion and limited vaccine efficacy.
There are two great opportunities to learn more about JanusMatrix and other tools for analyzing biologics and vaccines – in Washington DC at the World Vaccine Congress, and at the upcoming Westin Seminar on vaccines and biologics hosted by EpiVax in Seoul. See our events page for more information about these events. And for ‘hands on training’ using JanusMatrix, the University of Rhode Island Institute for Immunology and Informatics is offering a computational vaccinology seminar – BYOV! (build your own vaccine)! We hope to see you there!
*https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3739034/
1)Moise L, Gutierrez AH, Bailey-Kellogg C, Terry F, Leng Q, Abdel Hady KM, VerBerkmoes NC, Sztein MB, Losikoff PT, Martin WD, Rothman AL, De Groot AS. The two-faced T cell epitope: examining the host-microbe interface with JanusMatrix. Hum Vaccin Immunother. 2013 Jul;9(7):1577-86.
2)He L, De Groot AS, Gutierrez AH, Martin WD, Moise L and Bailey-Kellog C. Integrated assessment of predicted MHC binding and cross-conservation with self reveals patterns of viral camouflage. BMC Bioinf., 2014.
3)De Groot AS, Moise L, McMurry JA, Wambre E, Van Overtvelt L, Moingeon P, Scott DW, Martin W. Activation of natural regulatory T cells by IgG Fc-derived peptide “Tregitopes”. Blood. 2008 Oct 15;112(8):3303-11.
4)Moise L, Terry F, Gutierrez AH, Tassone R, Losikoff P, Gregory SH, Bailey-Kellogg C, Martin WD and De Groot AS. Smarter vaccine design will circumvent regulatory T cell-mediated evasion in chronic HIV and HCV infection. Front. Microbiol., 06 October 2014.
5)Losikoff PT, Mishra S, Terry F, Gutierrez A, Ardito MT, Fast L, Nevola M, Martin WD, Bailey-Kellogg C, De Groot AS, Gregory SH. HCV epitope, homologous to multiple human protein sequences, induces a regulatory T cell response in infected patients. J Hepatol. 2015 Jan;62(1):48-55.
